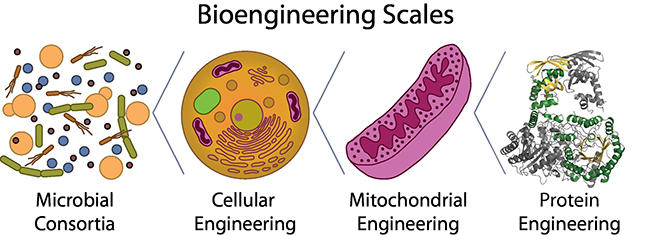
The mission of our group is to find biotechnological solutions to address challenges in sustainable energy, sustainable manufacturing, the environment, and human health. To achieve our goals, we integrate principles of microbiology, genetics, cell biology, biochemistry, biophysics, structural biology, and engineering. The length scales of our bioengineering efforts span microbial consortia, cells, organelles, and proteins (Figure 1).
Metabolic engineering
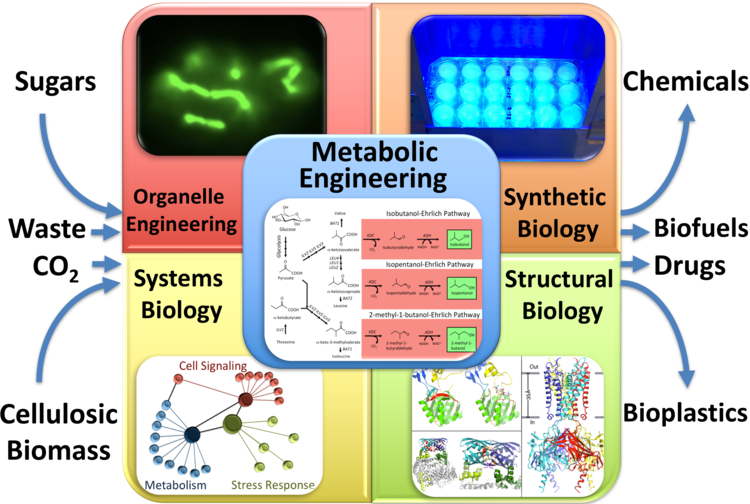
The core discipline in our lab is metabolic engineering. This is a rapid-moving field that aims to rewire the metabolism of microbes to make valuable products from renewable sources, such as sugars (including cellulosic biomass), waste, or even CO2; thus reducing the carbon footprint of manufacturing processes, and increasing their sustainability. Some of the products we are interested in include advanced biofuels (gasolines, diesels, and jet fuels), commodity chemicals (currently made from petroleum), specialty chemicals (including natural products for flavors, aromas, pigments, or pharmaceuticals), polymers (for biodegradable plastics), and proteins (for pharmaceuticals, enzymes, or materials).
Our efforts in metabolic engineering are boosted by other disciplines, which directly or indirectly assist in the implementation, control, and optimization of engineered metabolic pathways. These include organelle engineering, synthetic biology, systems biology, and structural biology/protein engineering (Figure 2).
Organelle/Mitochondrial Engineering
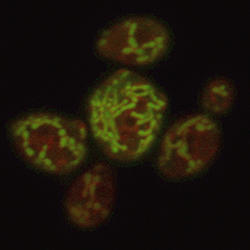
Subcellular engineering is an emerging and exciting field, in which metabolic pathways are targeted to specific organelles in the cell, in order to benefit from their unique environments, metabolites, and enzymes. In addition, the physical separation of organelles from the rest of the cell can confer pathway specificity, as well as prevent pathway toxicity, loss of intermediates, and byproduct formation. We are particularly interested in mitochondrial engineering. Our group demonstrated that targeting metabolic pathways to yeast mitochondria is an effective strategy to improve the production of advanced biofuels. By engineering mitochondrial physiology to control their size, number, and spatial distribution, it may be possible to further enhance metabolic pathways targeted to this highly dynamic, and versatile organelle (Figure 3).
Our group is also interested in developing completely de novo organelles with properties specifically designed for metabolic engineering applications. These synthetic organelles have the advantage of not being essential for normal cellular functions, and thus their exploitation is less likely to interfere with cell growth and health. In addition, these organelles can be designed to exclude all competing pathways, and thus enhance pathway specificity.
Synthetic Biology
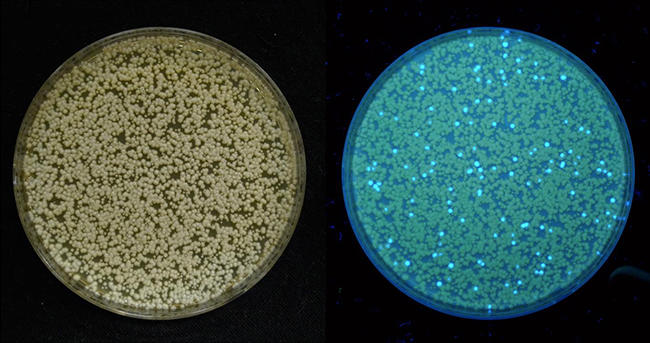
Synthetic biology has taken on different meanings to different researchers. In our group, synthetic biology implies the development of synthetic phenotypes beyond cellular production of chemicals. This involves designing and implementing biosensors, gene circuits, and optogenetic systems to enhance the screening, monitoring, control, and optimization of engineered metabolic pathways. Genetically encoded biosensors can be used to either monitor, screen, or select for a specific trait, for example production of a desired chemical (Figure 4). Such biosensors can enable high throughput technologies to accelerate strain development, and optimization. They can also be connected to gene circuits to perform logical operations in the cell, in order to confer automated dynamic control of metabolic pathways during fermentation.
Our group recently demonstrated that optogenetics can be used to dynamically control engineered microbes for chemical production. Optogenetics involves light-sensitive proteins, commonly transcription factors, whose activity responds to specific wavelengths of light. These proteins can be used to control genetic programs and metabolic pathways with light signals. Light enjoys most of the benefits of chemical inducers, with the advantage that it can be applied and removed instantly, without complex media changes, and independently of carbon source or nutrient composition. Thus, tunable and reversible light inputs can be used to optimize metabolic pathways, as well as to dynamically control fermentations, including in lab scale bioreactors (Figure 5). We have shown that optogenetics enables new strategies of periodic operation of fermentations, not only to control the transition from the growth phase to the production phase by switching light conditions, but also to control the fermentation during the production phase using periodic light pulses.
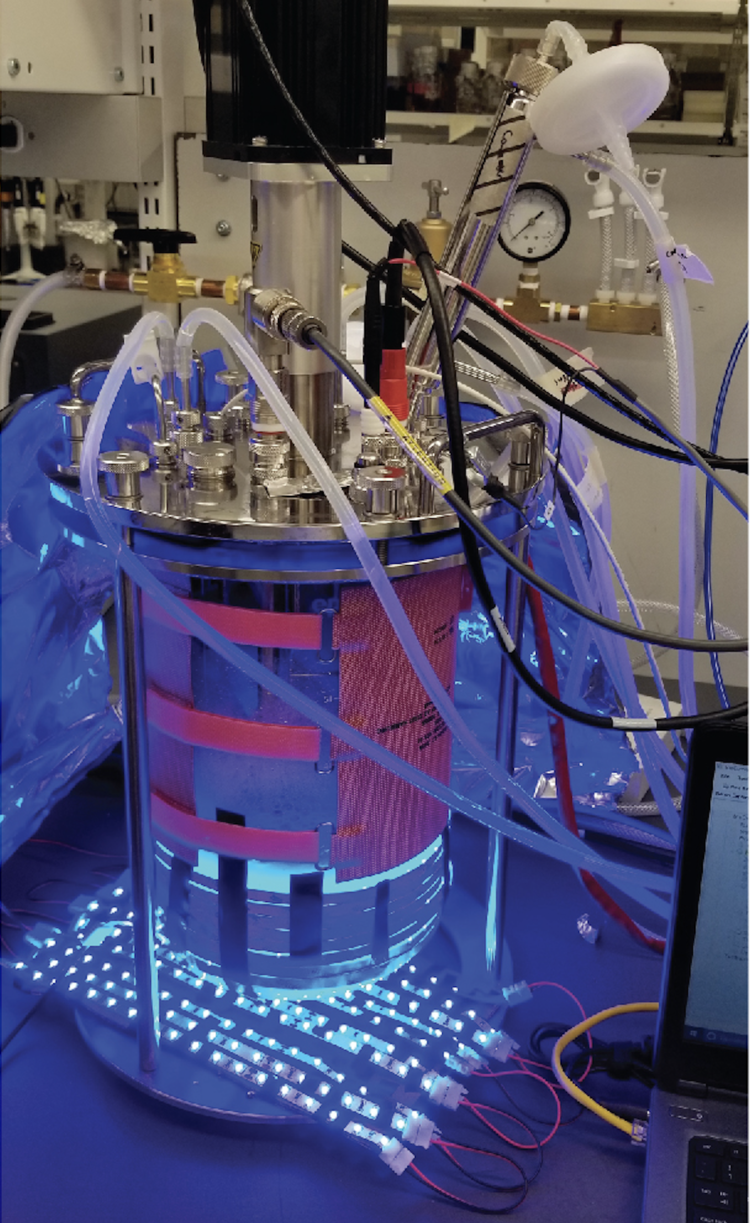

Figure 5. Light-controlled fermentation in a 5-L bioreactor.
Systems Biology
To achieve the levels of production of fuels and chemicals required for economic viability, it will be necessary to understand the effects that complex whole-cell biological systems have on their production, and take them into consideration when engineering strains. Similarly, it is important to understand the effects that engineered metabolic pathways have on normal physiology in order to develop robust strains that grow rapidly and show high tolerance to the products they make. Our strategy is to use various methods of detection, including biosensors, to run genome-wide screens to measure the effects of different perturbations on the production of molecules of interest, and vice versa. The data we obtain is useful to identify the cellular networks (metabolic, regulatory, signaling, etc.) involved in the production of fuels and chemicals, and the cellular tolerance of host organisms to these products.
Structural Biology and Protein Engineering
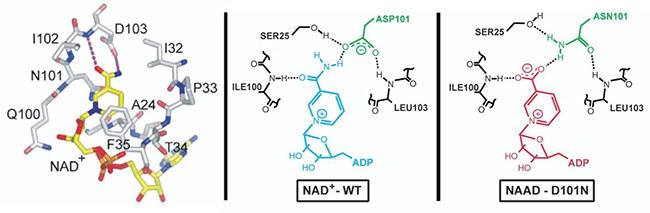
Our efforts in synthetic biology and metabolic engineering are complemented by fundamental studies on the molecular structure and function of the proteins involved, such as enzymes, transmembrane transporters, receptors, and transcription factors. To study these proteins in molecular detail we use different biophysical and biochemical methods, including X-ray crystallography. Understanding the relationship between the structure and function of these proteins significantly enhances our ability to do protein engineering in order to confer new functions of interest and relevance to metabolic engineering (Figure 6).